Asian longhorned beetle (ALB) is a wood-boring insect native to China and Korea. ALB attacks nearly all broadleaf trees in North America, but prefers native maples. Females lay eggs through the bark and larvae tunnel through the living tissue interrupting water and nutrient transport, killing the tree. ALB is easily transported in firewood, live trees or untreated lumber wood such as packing material used in shipping.
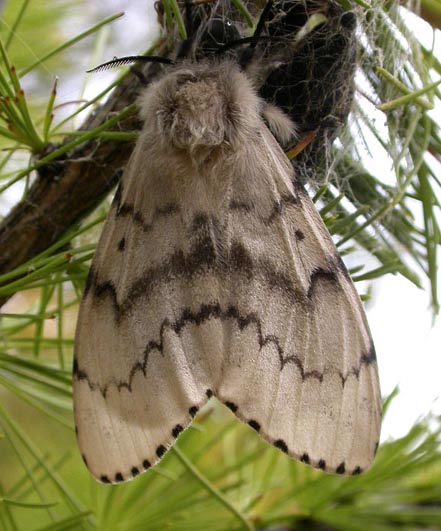
ALB was first introduced to North America in the 1990s, probably in solid wood packaging materials, such as crates or pallets. In 2003, ALB was discovered for the first time in Canada in an industrial park in the Toronto area. The infestation was controlled by removing host trees within a quarantine zone around the infestation to prevent further spread, at a cost of tens of millions of dollars over four years. Another ALB infestation was detected in Toronto in 2013. Eradication and detection efforts were initiated for a second time, and the area is being actively monitored to determine their success. Overall, more than US$500M has been spent on ALB eradication and associated monitoring in North America since the 1990s. [Picture courtesy of Brent Sinclair and Amanda Roe]
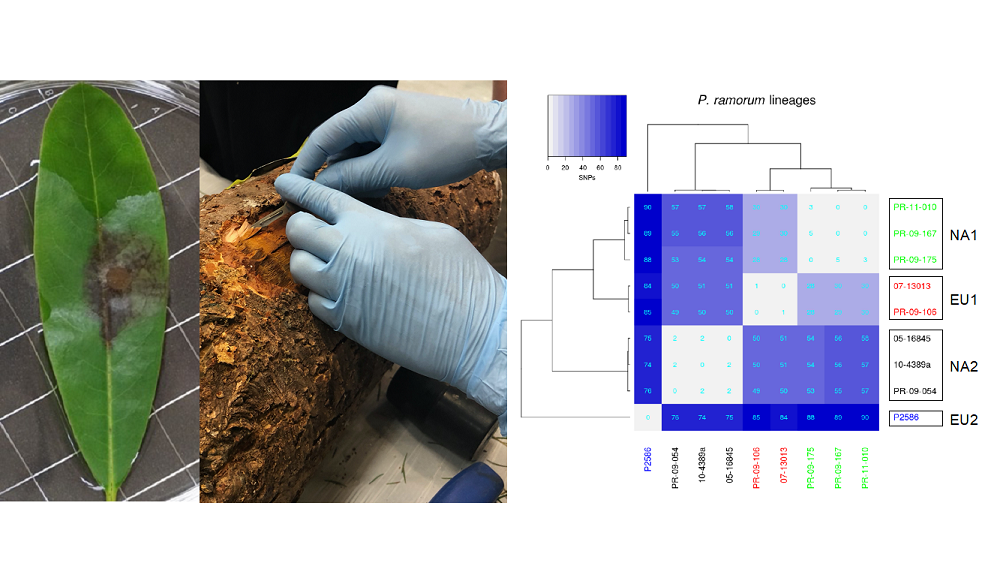
ALB have a broad native range in China, and we expect that some of the populations will be more likely to be able to overwinter in harsh Canadian winters. If we can identify those populations, it will allow resources to be prioritised towards the riskiest introduction events. One part of the BioSAFE team (led by Dr. Ilga Porth, Université Laval and Dr. Amanda Roe, Natural Resources Canada) is identifying genomic markers called single nucleotide polymorphisms (SNPs) to allow us to effectively identify the geographic source of a new infestation.
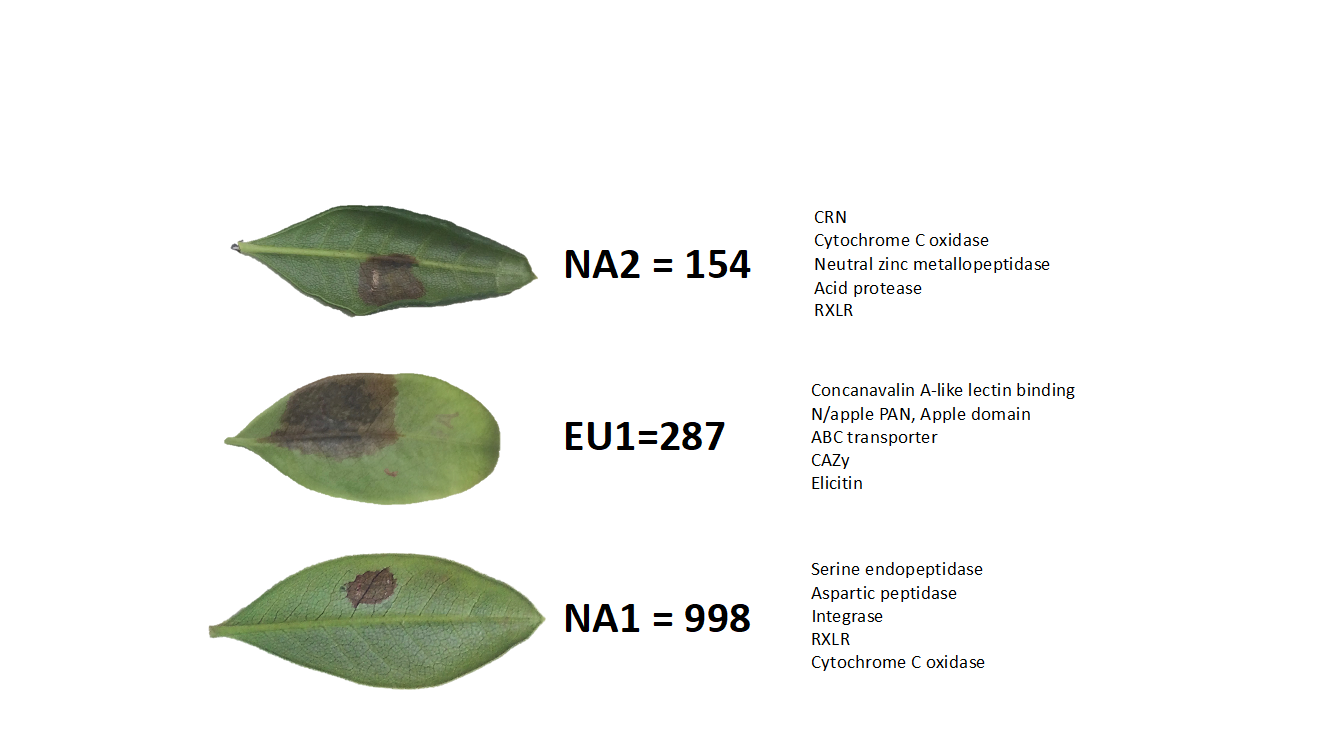
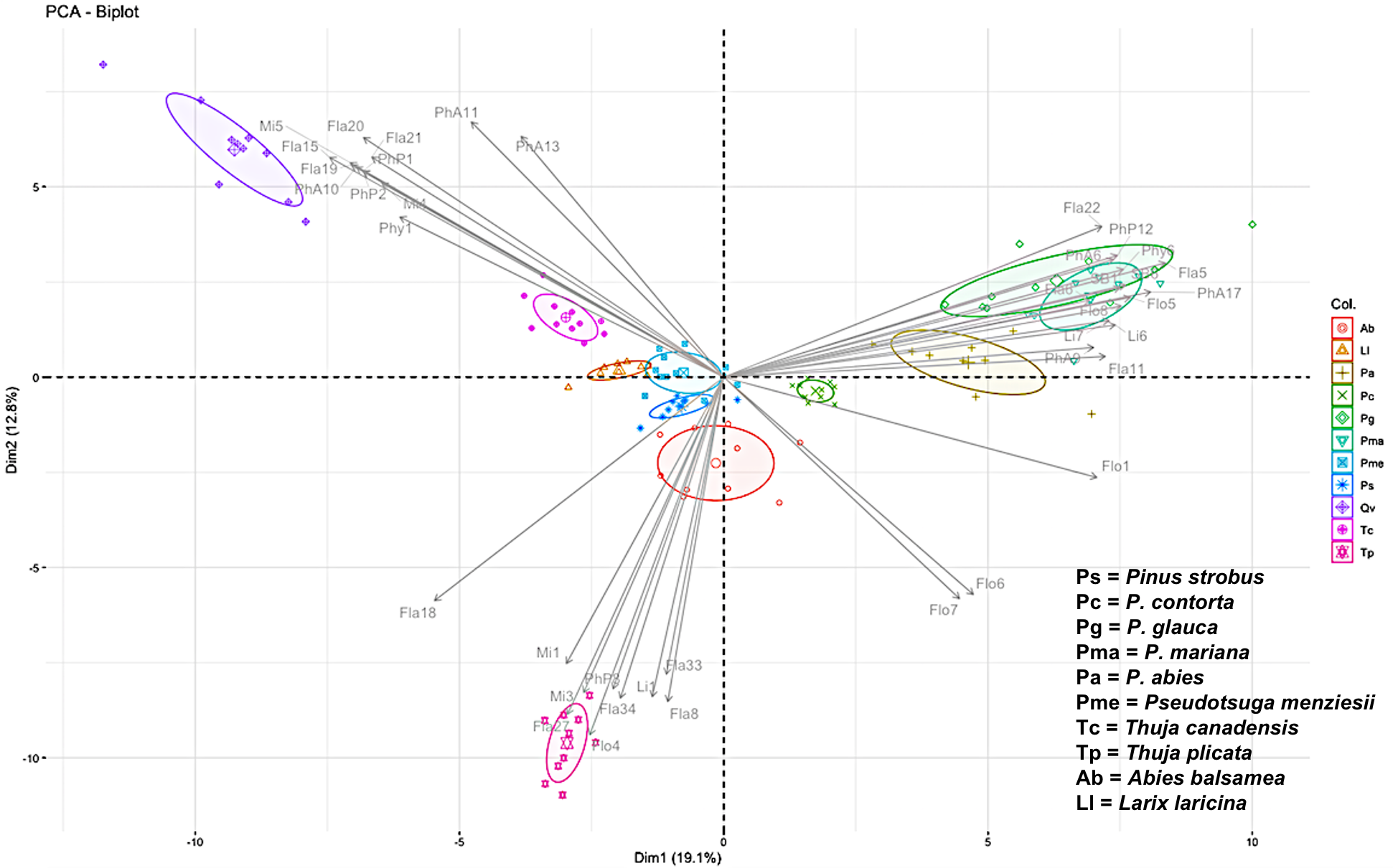
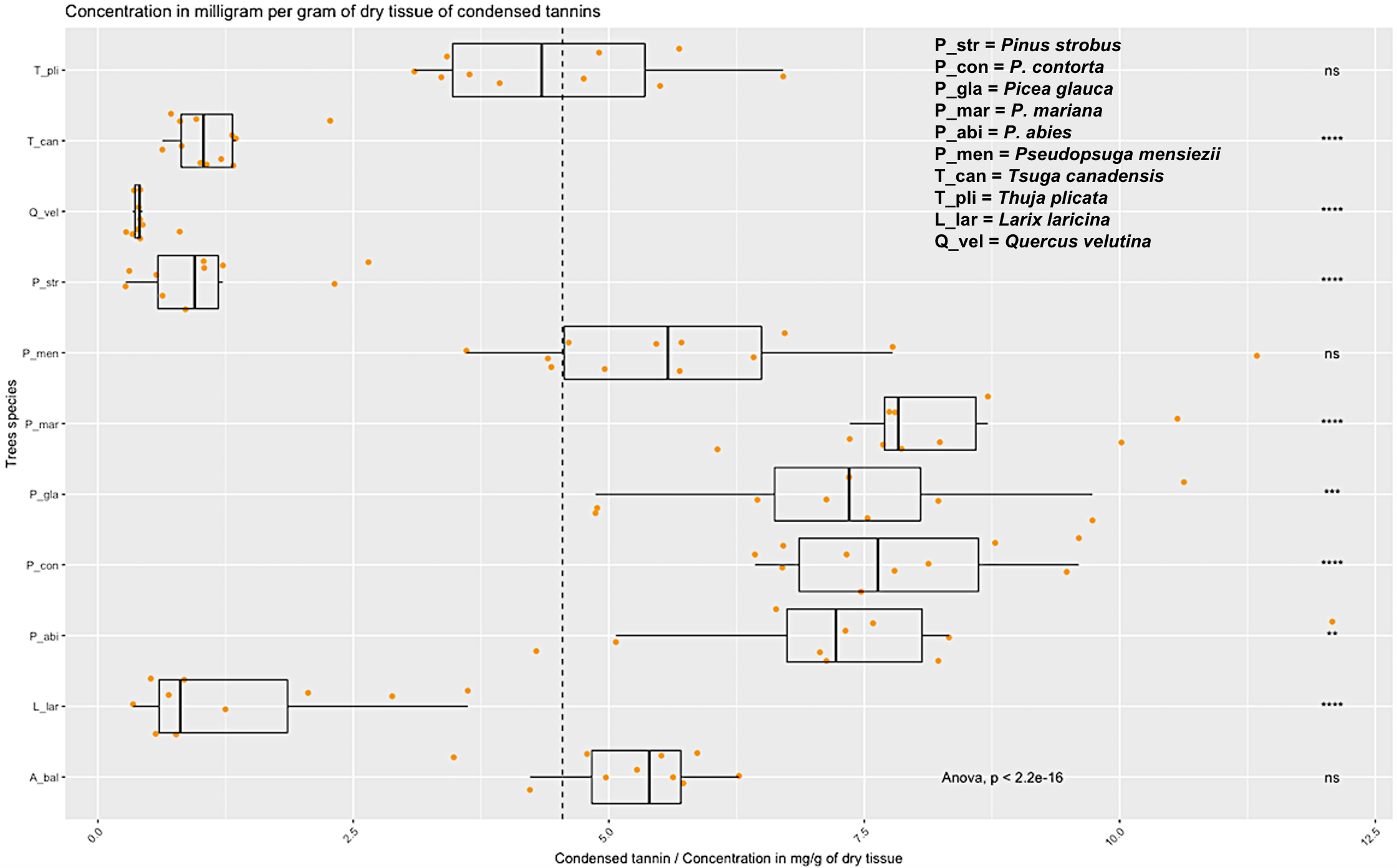
A parallel, related, approach is led by Dr. Brent Sinclair (Western University), Dr. Roe and Dr. Daniel Doucet (Natural Resources Canada), who aim to find genomic markers (Figures 1 and 2) that indicate the ability of a given ALB to survive winter. To do this, they are first determining the physiology responsible for overwintering success, and then using transcriptomics and metabolomics to identify the pathways and molecules that underlie those physiological mechanisms. They will then use the SNP database to identify SNPs that are likely to be speciically associated with enhanced overwintering capacity. This work is being performed at the natural resources Canada Insect Production and Quarantine Laboratory (IPQL) - the only Canadian facility certified to house ALB colonies.
Figure 1.

Figure 2

Flow charts by Dr. Alex Torson